Project Tasks16
Implement GEO Integration API
As a Backend Developer, I want to implement FastAPI endpoints to search and import healthy control samples from the NCBI GEO database so that Data Analysts can supplement patient data for comparative analysis. Endpoints: GET /geo/search?query=, POST /geo/import (by GEO series ID), GET /geo/datasets. Results stored in MySQL with integration status tracking.
Implement Theme & Global Styles
As a Frontend Developer, I want to implement the garnet-mg design system — color tokens (#9B111E primary, #50C878 accent, #0D0A0B bg), Merriweather/Open Sans typography, glassmorphism utilities, animated radial gradient root background, grain texture overlay, and reusable CSS custom properties — so that all pages inherit a consistent visual identity matching the mock designs. Remove any scaffold pages not present in the design (e.g. welcome page, dashboard/overview, dashboard/ai-assistant, dashboard/settings). This task must be completed before all page implementation tasks.
Implement Dataset Upload API
As a Backend Developer, I want to implement FastAPI endpoints for dataset upload (scRNAseq, scTCR, scBCR, Bulk TCR files) including multipart file upload support, file type validation (.h5ad, .loom, .csv, .tsv, .fastq.gz, .bam), 50GB per-file limit enforcement, metadata storage in MySQL, and file storage in MongoDB/object store. Endpoints: POST /datasets/upload, GET /datasets/{id}/status. Supports the Upload page and Upload Queue Panel.
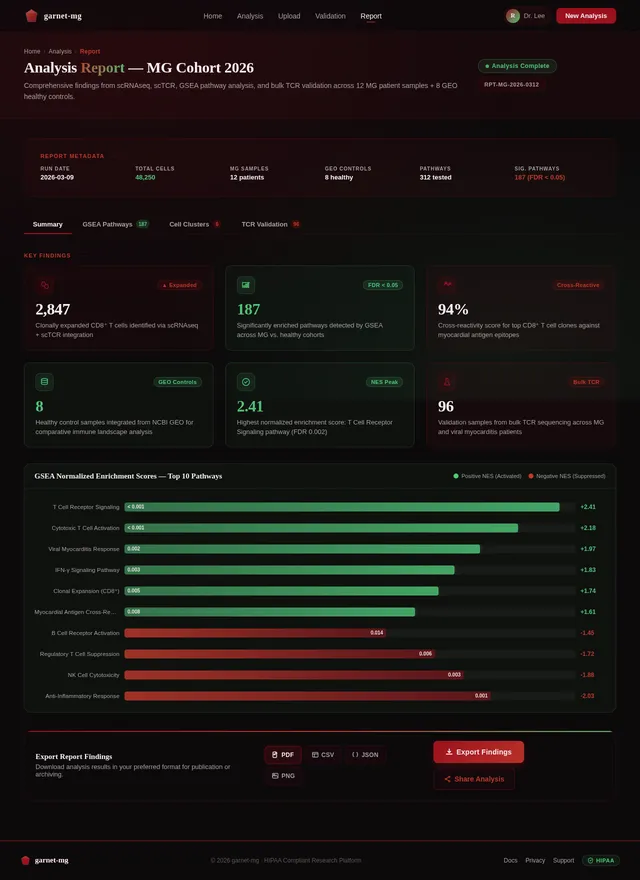
Build Report Page
As a Frontend Developer, I want to implement the Report page based on the existing v2 JSX design so that Researchers and Data Analysts can export and review their analysis findings. The page includes a structured report viewer showing GSEA results, TCR cross-reactivity scores, clonotype data, and export options (PDF, CSV, JSON). Navigates from Validation; export triggers file download.
Implement GSEA Analysis API
As a Backend Developer, I want to implement FastAPI endpoints to trigger and retrieve GSEA pathway analysis runs on uploaded scRNAseq/scTCR datasets so that Researchers can identify activated pathways in clonally expanded T cells. Endpoints: POST /analysis/gsea (params: gene set DB, threshold, permutations), GET /analysis/{run_id}/status, GET /analysis/{run_id}/results. Integrates with Langchain/LiteLLM for orchestration. Results include NES scores, p-values, FDR, enriched pathway lists.
Implement Bulk TCR Validation API
As a Backend Developer, I want to implement FastAPI endpoints to run the bulk TCR validation pipeline (Sample Input → TCR Repertoire Analysis → Cross-Reactivity Scoring → Myocardial Antigen Match) so that Researchers can validate cross-reactivity findings. Endpoints: POST /validation/run (cohort, antigen database, GEO toggle), GET /validation/{run_id}/status, GET /validation/{run_id}/results (CDR3 sequences, clonal frequency, similarity scores, confidence levels). Supports 500 MG + 100 viral myocarditis samples.
Build Data Integration Page
As a Frontend Developer, I want to implement the Data Integration page based on the existing v2 JSX design so that Data Analysts can import GEO samples and merge datasets for comparative analysis. The page includes: GEO sample import interface, dataset merge controls, QC metric previews, and integration status tracking. Navigates from Home; leads to Analysis page.
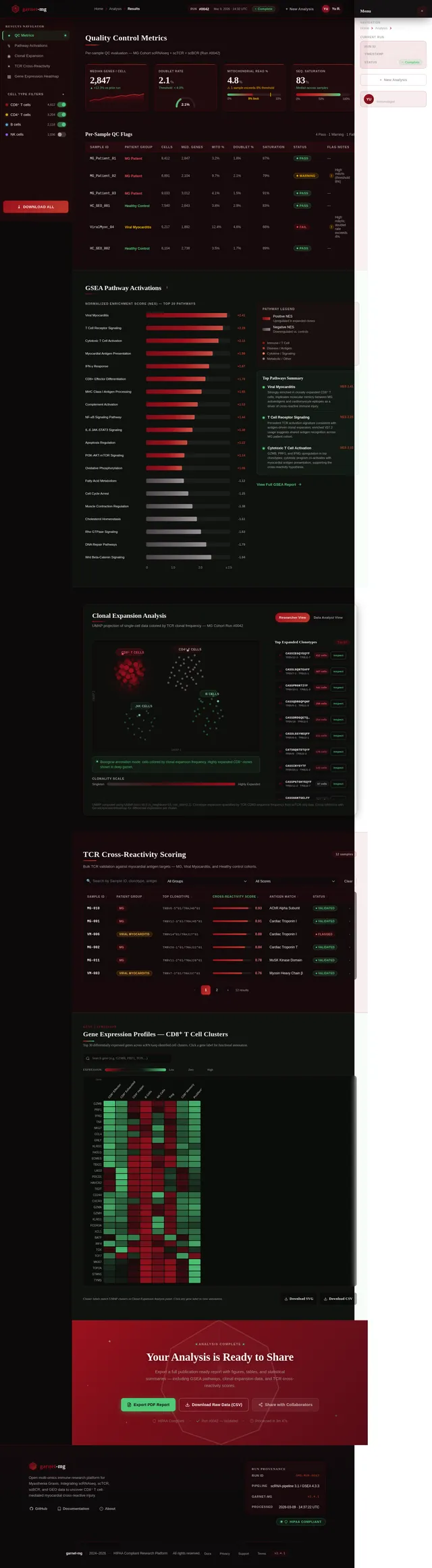
Build Results Page
As a Frontend Developer, I want to implement the Results page based on the existing v2 JSX design so that Researchers can view pathway activations and analysis outcomes. The page presents GSEA enrichment results, UMAP cluster visualizations, clonotype expansion data, and statistical summaries for the completed analysis run. Includes navigation to Validation page and Report export. Navigates from Analysis; leads to Validation and Report pages.
Implement Statistical Models API
As a Data Engineer, I want to implement backend endpoints for statistical comparison models (differential abundance testing, QC metrics computation, dataset merging/integration) so that Data Analysts can run comparative analyses between MG patient and GEO healthy control datasets. Endpoints: POST /analysis/statistical, GET /analysis/{run_id}/qc-metrics, POST /datasets/merge.
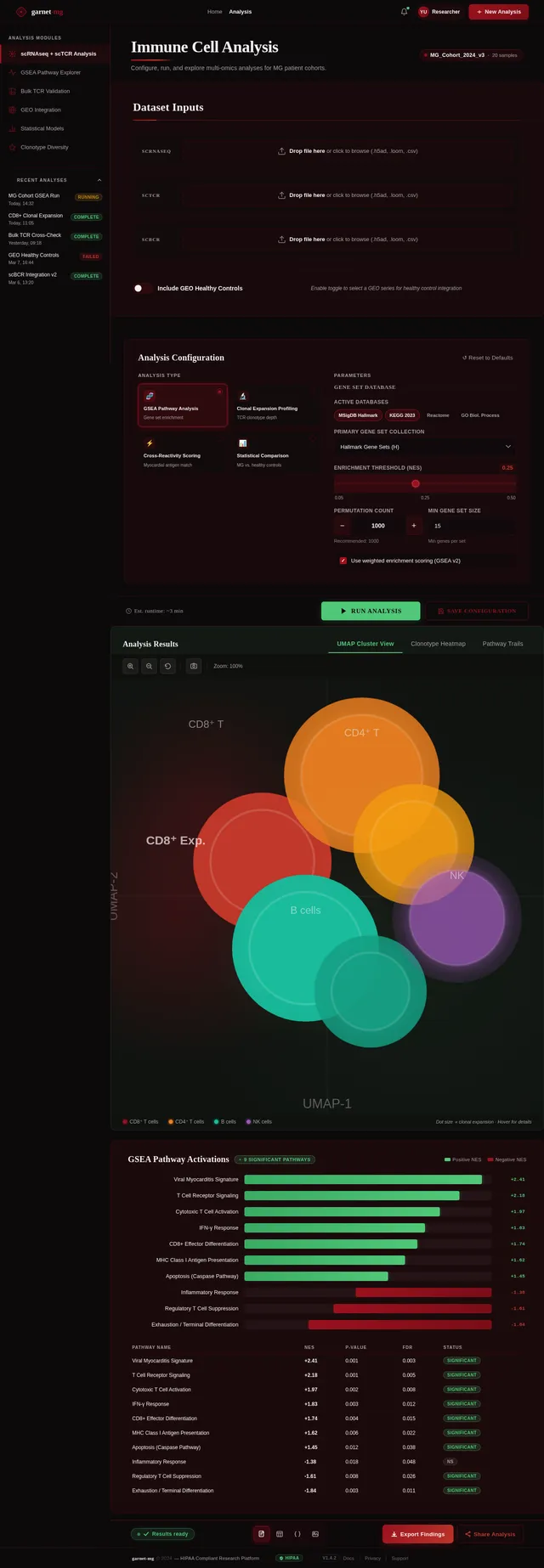
Build Analysis Page
As a Frontend Developer, I want to implement the Analysis page based on the existing v2 JSX design so that Researchers and Data Analysts can configure and run multi-omics analyses. The page includes: AnalysisTopBar (frosted-glass, breadcrumb, notification bell), AnalysisSidebar (module nav: scRNAseq+scTCR, GSEA, Bulk TCR, GEO Integration, Statistical Models, Clonotype Diversity; recent analyses list with status badges), AnalysisPageHeader (title, subtitle, dataset status chip), DatasetInputPanel (scRNAseq/scTCR/scBCR dropzones, GEO toggle), AnalysisConfigPanel (analysis type radio cards, dynamic parameter form), RunAnalysisCTA (estimated runtime, Run Analysis button with progress state, Save Configuration), ResultsVisualizationPanel (UMAP/Heatmap/Pathway tabs, interactive scatter plot, cluster tooltips), GSEAPathwayOutput (ranked bar chart, pathway detail drawer, data table), AnalysisExportBar (format icons, Export Findings button), AnalysisFooter. Navigates from Upload; leads to Results.
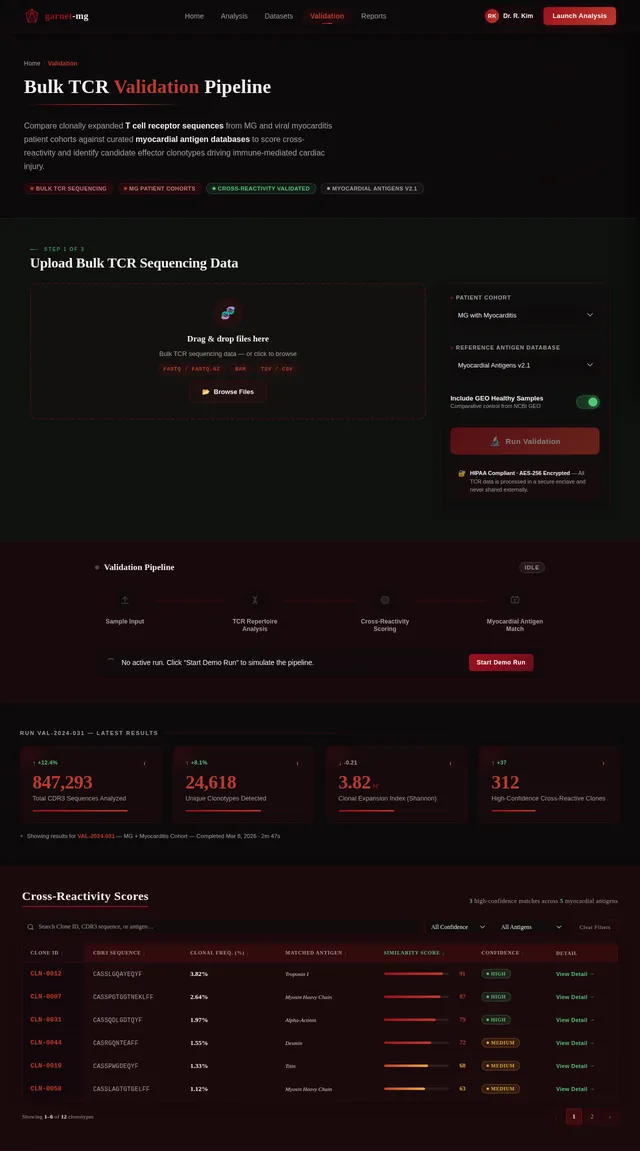
Build Validation Page
As a Frontend Developer, I want to implement the Validation page based on the existing v2 JSX design so that Researchers can compare clonally expanded TCR sequences from MG patients against myocardial antigen databases to score cross-reactivity. The page includes: NavBar (Validation active state), ValidationPageHero (breadcrumb, Merriweather headline 'Bulk TCR Validation Pipeline', garnet rule, subheadline, tag pills, gem watermark), ValidationUploadPanel (drag-and-drop for FASTQ/BAM/TSV, cohort dropdown, antigen database selector, GEO toggle, Run Validation CTA), PipelineStatusTracker (4-stage animated pipeline), TCRRepertoireStats (4 metric cards with animated counters), CrossReactivityScoreTable (sortable/filterable table with CDR3/antigen/score/confidence columns), ClonotypeDiversityChart (stacked bar + radar chart), MyocardialAntigenMatchPanel (antigen heatmap + top-5 summary), ValidationRunHistory (card list with status badges), ExportReportCTA, Footer. Navigates from Results; leads to Report.
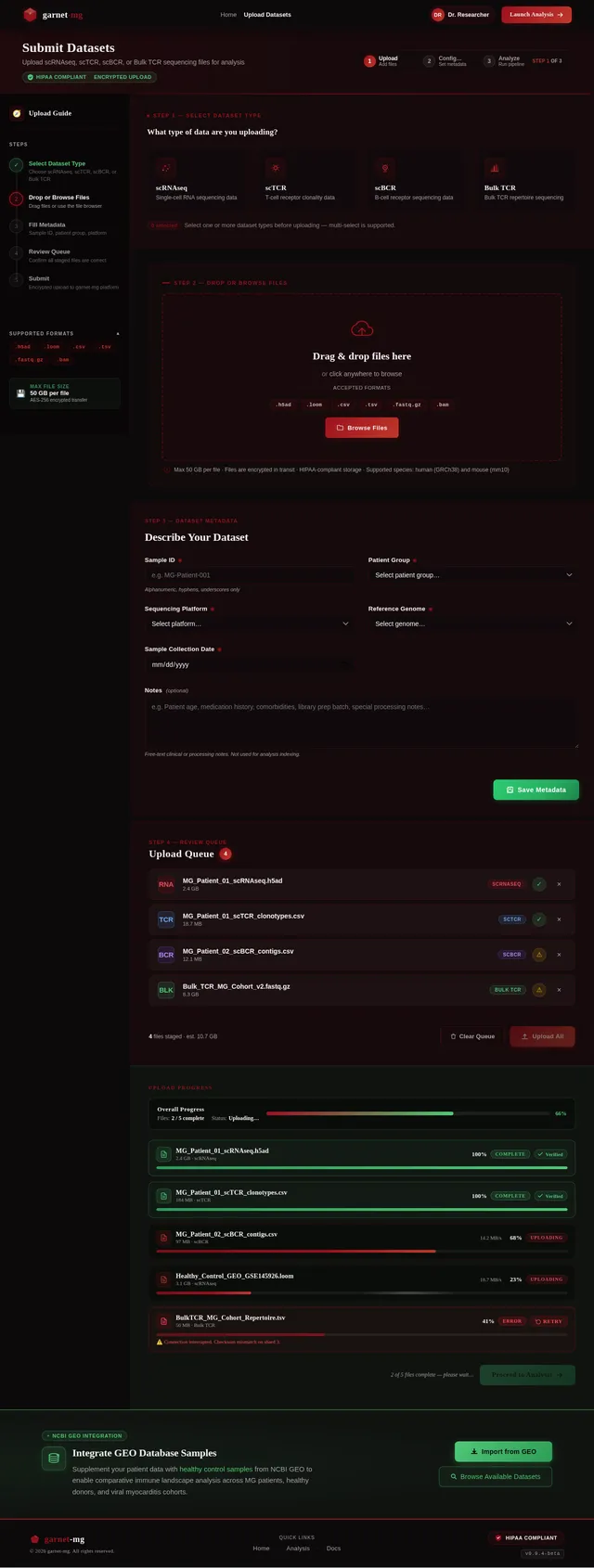
Build Upload Page
As a Frontend Developer, I want to implement the Upload page based on the existing v2 JSX design so that Researchers can submit scRNAseq, scTCR, scBCR, and Bulk TCR sequencing datasets. The page includes: TopBar (frosted-glass, breadcrumb), UploadPageHeader (page title, HIPAA chip, 3-step progress indicator), UploadSidebar (upload guide checklist, supported formats, max file size), DatasetTypeSelector (4 toggleable cards), FileDropZone (drag-and-drop with garnet dashed border), MetadataForm (2-column grid: Sample ID, Patient Group, Platform, Reference Genome, Date, Notes), UploadQueuePanel (staged files list with metadata completeness), UploadProgressTracker (per-file progress bars, garnet-to-emerald on complete), GEOIntegrationBanner, Footer. On mobile: sidebar collapses to horizontal step strip. Navigates from Home; leads to Analysis page.
Implement Clonotype Analysis API
As a Backend Developer, I want to implement FastAPI endpoints for clonotype diversity analysis (Shannon entropy, Simpson index, Chao1, ACE, Observed clones, Coverage) and clonal expansion profiling across patient cohorts (MG+myocarditis, MG only, Healthy Control) so that Researchers can quantify immune repertoire dynamics. Endpoints: POST /analysis/clonotype, GET /analysis/{run_id}/diversity-metrics, GET /analysis/{run_id}/expansion-profile.
Build Home Page
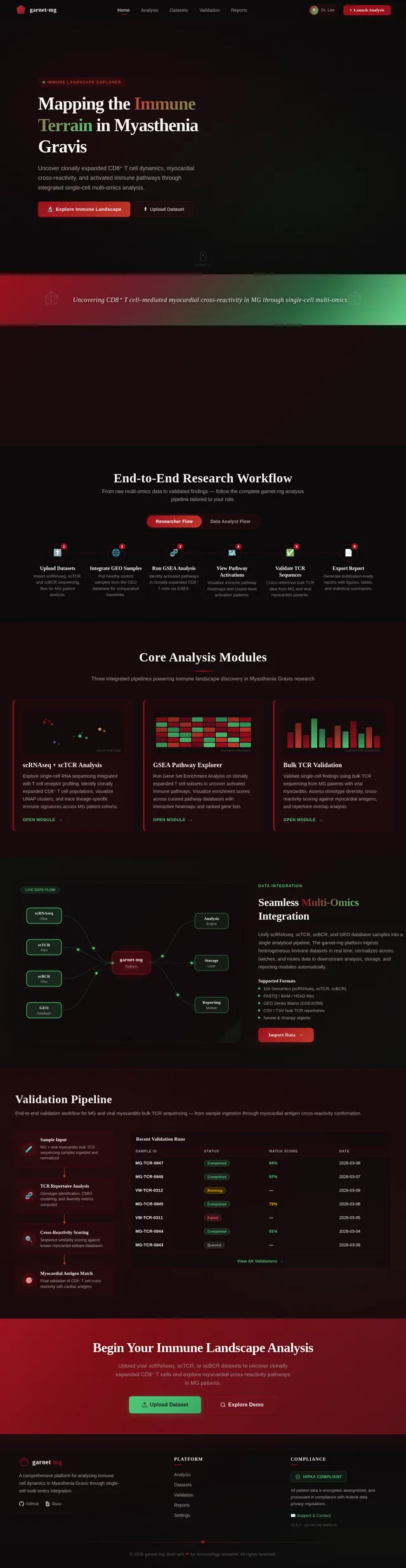
As a Frontend Developer, I want to implement the Home page based on the existing v2 JSX design so that Researchers can explore the interactive 3D immune landscape visualization. The page includes: NavBar (frosted-glass sticky), Hero (Three.js/WebGL 3D immune terrain with CD8+/CD4+/B/NK cell clusters, garnet-to-emerald gradient transitions, cluster tooltips), ResearchMissionBanner, ImmuneDataHighlights (stats row with animated counters), WorkflowOverview (animated stepper), FeaturedAnalysisModules (3-column card grid), DataIntegrationVisual (animated data-flow diagram), ValidationPipeline (pipeline diagram + validation runs table), QuickAccessCTA, and Footer. On mobile: 3D canvas degrades to static image with CSS animation. Links to: Upload page, Analysis page.
Implement Report Export API
As a Backend Developer, I want to implement FastAPI endpoints to generate and export analysis reports in PDF, CSV, JSON, and PNG formats so that Researchers can download and share their validation findings. Endpoints: POST /reports/generate (analysis_run_id, format), GET /reports/{report_id}/download. PDF includes GSEA results, TCR cross-reactivity scores, clonotype data, and charts.
Integrate AI Hypothesis Generation
As an AI Engineer, I want to integrate GPT-based LLM (via LiteLLM routing and Langchain) to provide AI-assisted hypothesis generation and user-friendly result interpretation on the Analysis and Results pages so that Researchers can receive contextual insights about their immune landscape findings. Implement: POST /ai/interpret (analysis_run_id), streaming response endpoint, prompt templates for GSEA/TCR cross-reactivity context.


No comments yet. Be the first!