System Requirements
System Requirements Document (SRD)
Project Name: garnet-mg
1. Introduction
The garnet-mg project aims to explore the mechanisms underlying immune-mediated damage in Myasthenia Gravis (MG) patients, specifically focusing on the hypothesis that clonally expanded CD8⁺ T cells mediate myocardial cross-reactive immune injury. This research builds upon prior findings using scRNAseq, scTCR, and scBCR sequencing, as well as GEO database integration of healthy samples. The project will leverage cutting-edge technologies and data validation methods to uncover novel insights into MG pathogenesis.
This document outlines the system requirements for the garnet-mg project, ensuring alignment with the research goals and providing a clear roadmap for implementation.
2. System Overview
The garnet-mg system will serve as a comprehensive platform for analyzing immune cell dynamics in MG patients. It will integrate multi-omics data, including single-cell sequencing and bulk TCR sequencing, to identify clonally expanded T cells and their cross-reactivity with myocardial antigens. The system will also facilitate hypothesis validation through mechanism experiments and advanced computational modeling.
Key objectives include:
- Data integration from multiple sources (scRNAseq, scTCR, scBCR, GEO database).
- Identification of clonally expanded CD8⁺ T cells and their functional pathways.
- Validation of findings using bulk TCR sequencing data from MG patients, including those with viral myocarditis.
- Mechanistic exploration of immune-mediated myocardial injury.
3. Functional Requirements
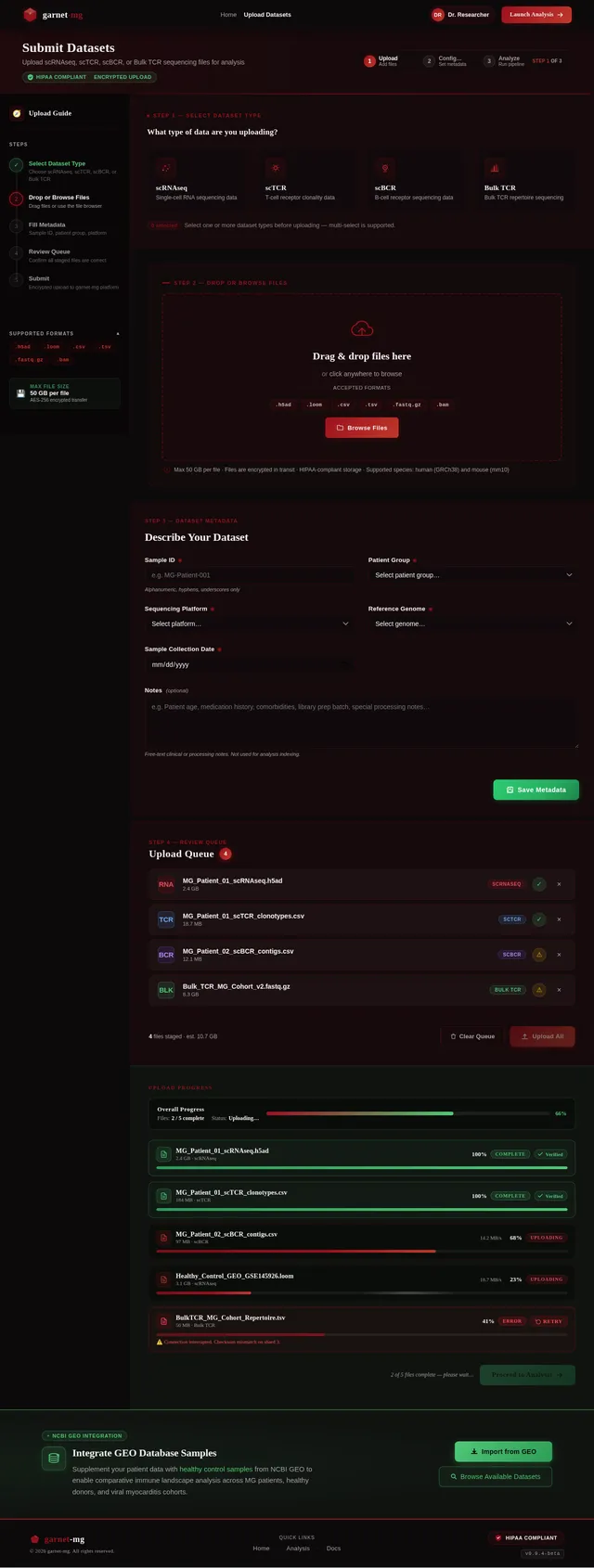
- As a Researcher, I should be able to upload scRNAseq, scTCR, and scBCR datasets for analysis.
- As a Researcher, I should be able to integrate GEO database healthy sample data for comparative analysis.
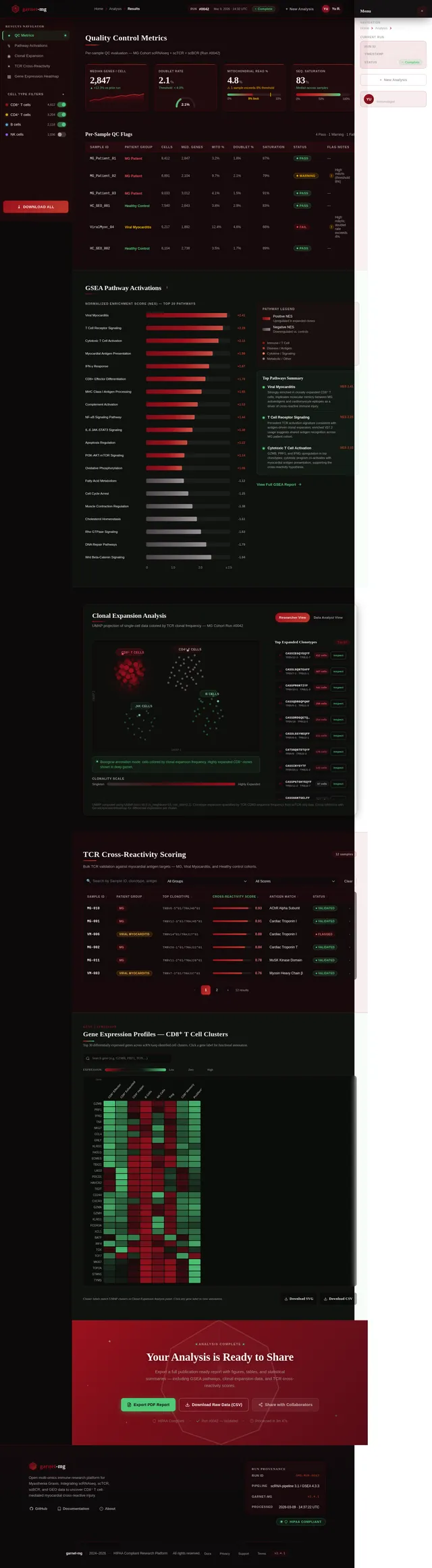
- As a Researcher, I should be able to perform GSEA analysis to identify activated pathways in clonally expanded T cells.
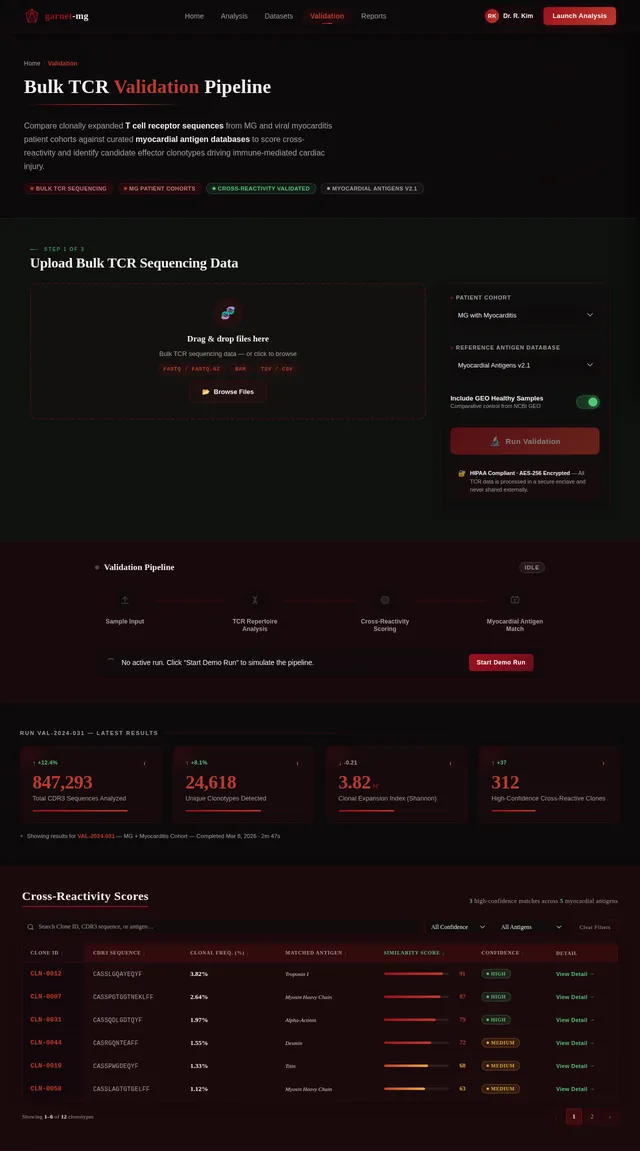
- As a Researcher, I should be able to validate findings using bulk TCR sequencing data from MG patients.
- As a Researcher, I should be able to conduct mechanism experiments to test the hypothesis of myocardial cross-reactivity.
4. User Personas
4.1 Researcher
- Description: Immunologists, bioinformaticians, and clinicians working on MG and related autoimmune diseases.
- Goals: To uncover mechanisms of immune-mediated damage in MG patients and validate findings through experiments.
- Skills: Expertise in immunology, bioinformatics, and single-cell sequencing technologies.
4.2 Data Analyst
- Description: Specialists in data integration and statistical analysis.
- Goals: To ensure accurate and meaningful analysis of multi-omics datasets.
- Skills: Proficiency in data processing tools and statistical modeling.
5. Visuals Colors and Theme
The garnet-mg project will adopt a theme inspired by the garnet gemstone, symbolizing clarity and precision.
Color Palette:
- Primary: Deep Garnet Red (#9B111E) — representing passion and focus.
- Secondary: Soft Gray (#D3D3D3) — for neutral backgrounds and contrast.
- Accent: Emerald Green (#50C878) — symbolizing growth and discovery.
Typography:
- Headings: Bold Serif (e.g., Merriweather).
- Body Text: Clean Sans-Serif (e.g., Open Sans).
Visual Elements:
- Gradient backgrounds transitioning from garnet red to emerald green.
- Subtle animations highlighting data insights and transitions.
6. Signature Design Concept
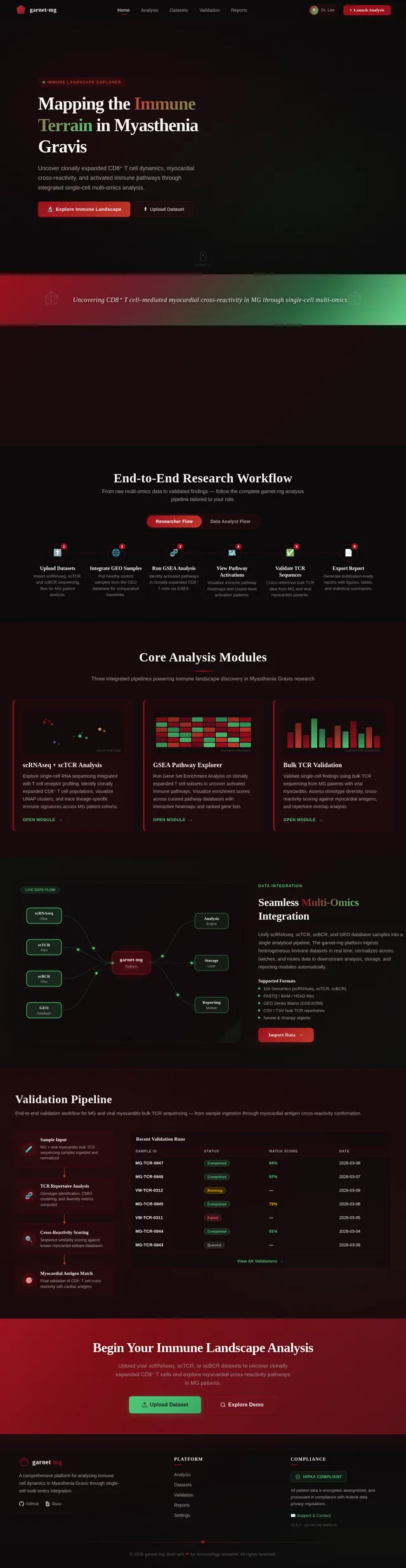
Interactive Immune Landscape Visualization
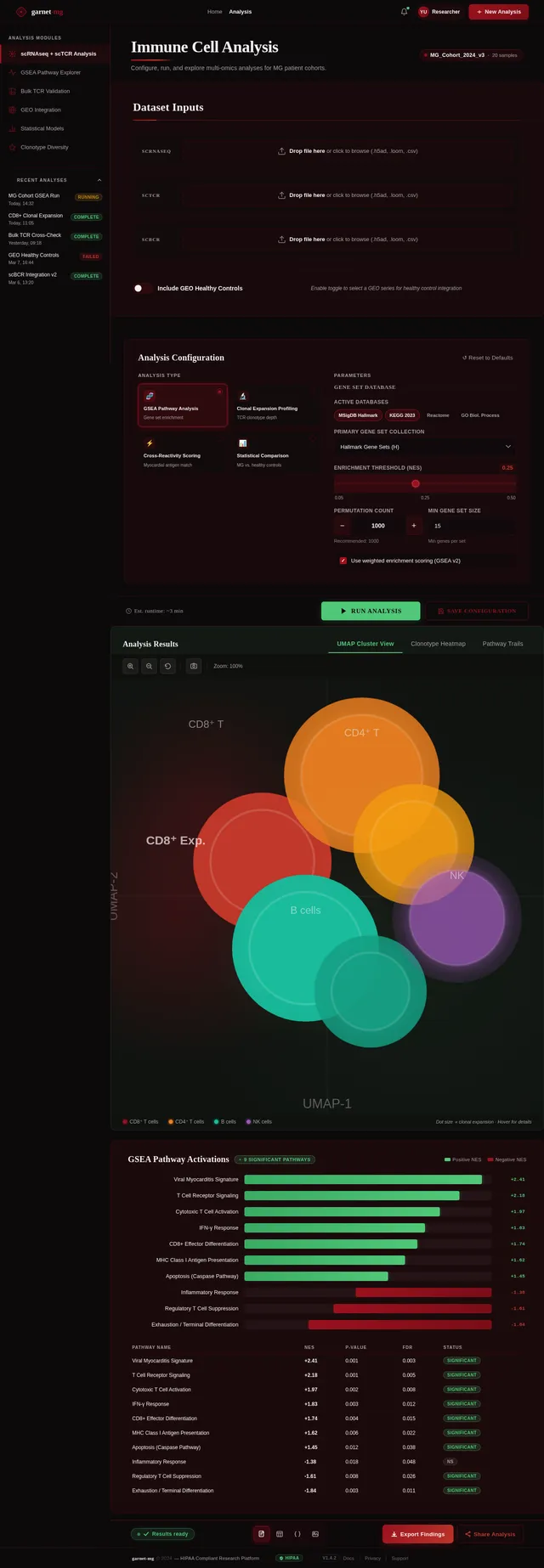
The homepage of garnet-mg will feature a 3D interactive immune landscape. Users will navigate through a virtual "immune terrain" where each cell type (e.g., CD8⁺ T cells, CD4⁺ T cells, B cells) is represented as dynamic clusters on a floating 3D map.
Key Features:
- Dynamic Clusters: Each cluster pulsates and changes size based on real-time data inputs (e.g., clonally expanded cells grow larger).
- Pathway Activation Trails: Activated pathways will light up as glowing trails connecting clusters, visually representing GSEA results.
- User Interaction: Clicking on a cluster zooms into detailed data, including gene expression profiles, clonality metrics, and pathway activations.
- Color Shifts: The background subtly transitions between garnet red and emerald green based on the user's navigation, symbolizing discovery and clarity.
- Micro-Interactions: Hovering over elements triggers tooltips with concise explanations and links to deeper insights.
This immersive design will captivate users, making the research journey both intuitive and visually engaging.
7. Non-Functional Requirements
- Performance: The system must process large multi-omics datasets efficiently, with results generated within 5 minutes for standard analyses.
- Scalability: Support for increasing data volumes as new samples are added.
- Security: Ensure data privacy and compliance with HIPAA regulations for patient data.
- Usability: Provide an intuitive interface for researchers with varying levels of technical expertise.
8. Tech Stack
Frontend:
- React for Web (to create an interactive and responsive user interface).
Backend:
- Python with FastAPI (for efficient data processing and API management).
Database:
- MySQL or MariaDB (for structured data storage and alembic migrations).
- MongoDB (for unstructured data storage).
AI Models:
- GPT 5.2 (for user-friendly responses and hypothesis generation).
- Google Nano Banana (for image generation in visualizations).
AI Tools:
- Langchain (for advanced data analysis workflows).
- Litellm (for LLM routing).
Orchestration:
- Docker and docker-compose (for local development).
- Kubernetes (for server-side orchestration).
9. Assumptions and Constraints
Assumptions:
- Researchers have access to scRNAseq, scTCR, and scBCR datasets.
- Bulk TCR sequencing data includes sufficient samples for validation.
- GEO database integration is feasible and provides relevant healthy sample data.
Constraints:
- Computational resources must support large-scale data analysis.
- Data privacy regulations must be strictly adhered to.
10. Glossary
- MG: Myasthenia Gravis, an autoimmune disease affecting neuromuscular function.
- scRNAseq: Single-cell RNA sequencing, a technique for analyzing gene expression at the single-cell level.
- scTCR: Single-cell T-cell receptor sequencing, used to study T-cell clonality.
- scBCR: Single-cell B-cell receptor sequencing, used to study B-cell clonality.
- GSEA: Gene Set Enrichment Analysis, a method for identifying activated pathways.
- CD8⁺ T Cells: A subtype of T cells involved in immune responses, often cytotoxic.
- Bulk TCR Sequencing: A method for analyzing T-cell receptor diversity in bulk samples.
宇,我希望这份SRD能够清晰地传达你的研究目标和技术需求,同时为项目的实施提供明确的指导!如果有任何需要调整的地方,请随时告诉我!


No comments yet. Be the first!