Project Tasks11
Implement Theme & Structure from Mock-Design
As a Frontend Developer, I want to implement the visual theme and structural layout from all mock-design pages so that the scaffold project pages look exactly identical to the designed pages. This includes applying the green (#16A34A), purple (#7C3AED), and blue (#1D4ED8) accent color system, typography (Arial/Calibri), glassmorphism effects (backdrop-filter blur, rgba backgrounds), SlideChrome and FooterBar shared components, AccentDivider gradient bars, and removing any pages not required by the SRD. This task must be completed independently before any other page-specific tasks begin.
Implement Color & Theme Alignment
As a Frontend Developer, I want to perform a full color and theme alignment audit across all slide pages to ensure consistent application of: green (#16A34A), purple (#7C3AED), and blue (#1D4ED8) accent colors, Arial/Calibri typography with correct weight hierarchy, glassmorphism card styles (backdrop-filter blur, rgba backgrounds, border-radius), accent divider bars (green→purple→blue gradient), SlideChrome and FooterBar consistency, and responsive breakpoints (320px mobile, 768px tablet, 1024px desktop). Fix any visual inconsistencies found across all 12 slide pages.
Build Title Slide Page
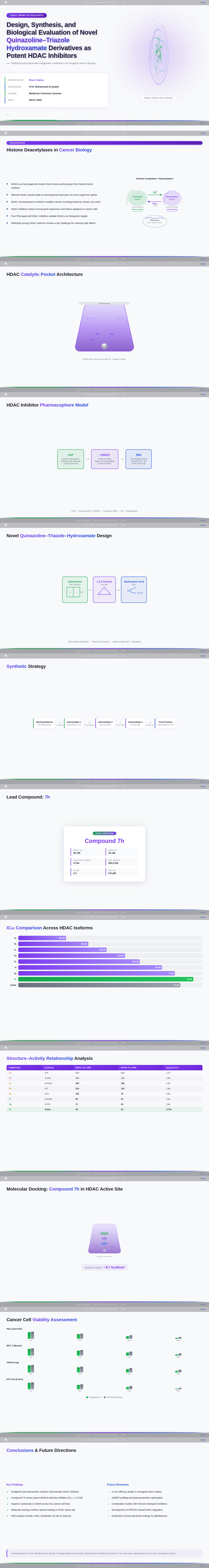
As a Frontend Developer, I want to implement the Title Slide page based on the existing JSX design (v3). The page features: dynamic 3D rotating HDAC enzyme molecular visualization with Zn²⁺ silver metallic sphere, gradient background blending green/purple/blue, centered bold title 'Design, Synthesis, and Biological Evaluation of Novel Quinazoline–Triazole Hydroxamate Derivatives as Potent HDAC Inhibitors', student names, university name, course name, professor name, and date. Includes SlideChrome top bar, TitleBlock, PresentationMeta, InstitutionBadge, MolecularVisualization, AccentDivider, and FooterBar sections. This is the entry point for both Audience and Presenter user flows.
Build Introduction Slide Page
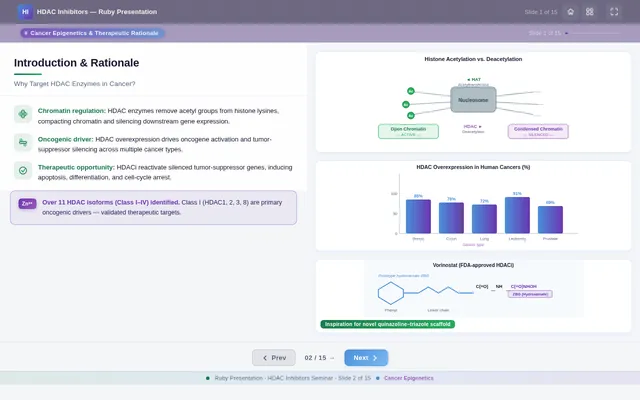
As a Frontend Developer, I want to implement the Introduction Slide page based on the existing JSX design (v4). The page covers Cancer Epigenetics & Therapeutic Rationale. Left column: SlideHeadline with title 'Cancer Epigenetics & Therapeutic Rationale', RationaleText card with 4 bullet points (cancer epigenetics, histone acetylation, HDAC isoforms 1/2/3/6/8, Vorinostat limitations), KeyInsightCallout with structural optimization rationale. Right column: EpigeneticsVisual (histone acetylation vs deacetylation diagram), HDACOverexpressionVisual (pathway diagram), VorinostatStructure (SAHA chemical structure with IC₅₀ ≈ 0.160 µM). Includes TopBar, ContextBanner, SlideNavigation, and BottomBar. This is step 2 in the Audience user flow — Read Rationale.
Build Slides Overview Page
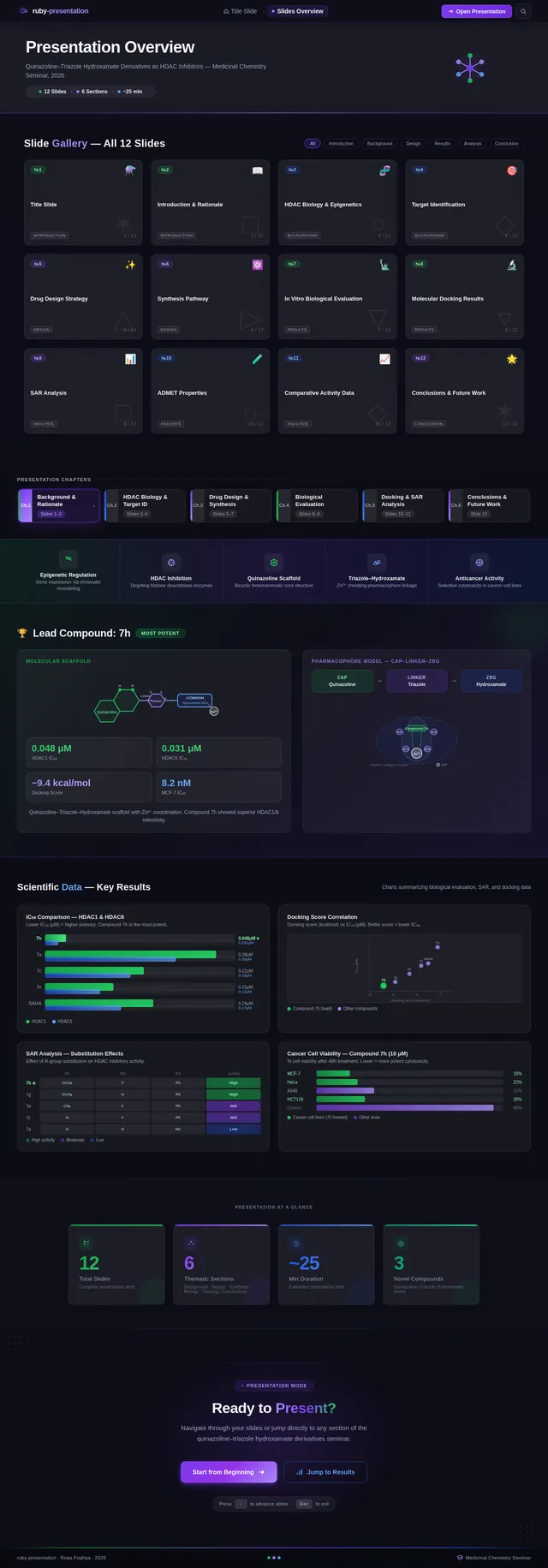
As a Frontend Developer, I want to implement the Slides Overview page based on the existing JSX design (v4). The page includes: OverviewNavBar with ruby-presentation wordmark, OverviewHero banner showing 'Presentation Overview' with animated molecule icon and slide count indicator (12 Slides · 6 Sections · ~25 min), SlideGridGallery with responsive 4/3/2 column thumbnail grid for all 12 slides, SectionChapters horizontal chapter navigator, KeyThemesBar with 5 scientific themes, PresentationStats with 4 stat cards, NavigationCTA, and OverviewFooter. Dark navy animated background with molecular lattice overlay. This is step 2 in the Presenter user flow — Review Structure.
Build Content Slide Page
As a Frontend Developer, I want to implement the Content Slide page based on the existing JSX design (v3). The page presents structured research content. Left column (55%): ContentOutlinePanel with frosted-glass bullet rows covering HDAC mechanisms, Class I/II overexpression in cancers, Zn²⁺ catalytic pocket inhibition target, quinazoline–triazole scaffold dual benefit, and nanomolar IC₅₀ values; KeyConceptsGrid with 2×2 concept cards (Epigenetic Regulation, Histone Acetylation, Class I HDACs, Zn²⁺ Catalysis). Right column (45%): ScientificDiagramPanel (histone acetylation/deacetylation SVG diagram), DataHighlightCard ('>80% solid tumors' stat with progress visualization). Includes SlideChrome, SlideTitleBar, ReferencesStrip, and FooterBar. This is step 3 in the Presenter user flow — Present Research.
Build Science Slide Page
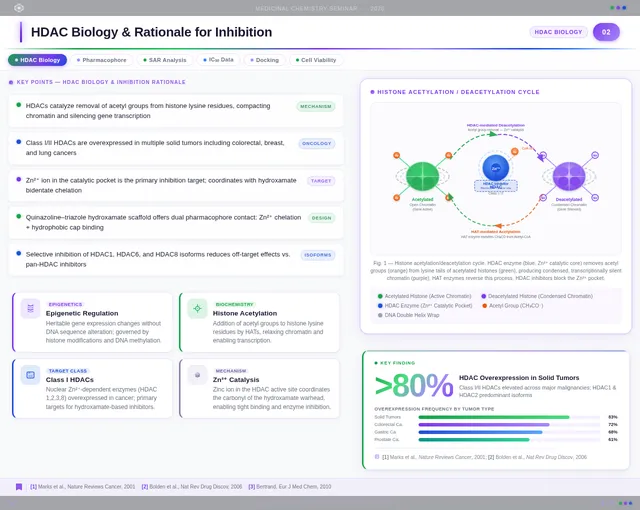
As a Frontend Developer, I want to implement the Science Slide page based on the existing JSX design (v4). Topic: 'Scientific Background & Mechanism of Action'. Includes SlideChrome, SlideTitleBar with purple left-border accent and 'HDAC Biology' category pill badge, ConceptIntroRow with three concept chips (Epigenetic Regulation, HDAC Enzyme Inhibition, Zn²⁺ Catalytic Mechanism), HDACMechanismDiagram (deacetylation cycle with Zn²⁺ sphere, pulsing silver shimmer animation, His142/His143/Tyr306 residue labels), HistoneAcetylationPanel (two-state chromatin diagram), MolecularDockingPanel (PyMOL-style docking visualization with IC₅₀ = 0.03 µM annotation), PharmacophoreAnnotation (quinazoline–triazole–hydroxamate 2D structure with SAR table), KeyTakeawayBar, and FooterBar. This is step 3 in the Audience user flow — View Diagrams.
Build Diagrams Slide Page
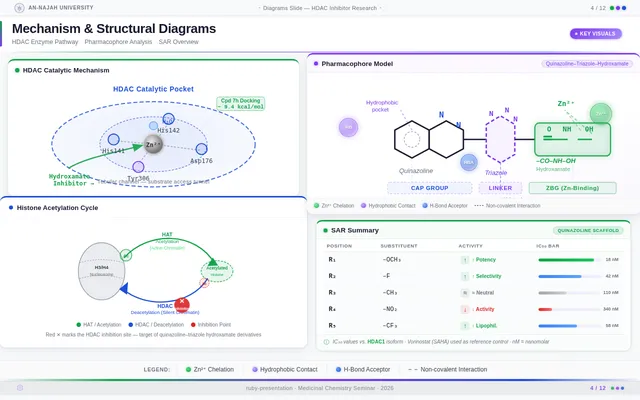
As a Frontend Developer, I want to implement the Diagrams Slide page based on the existing JSX design (v4). The page displays high-quality scientific visualizations including: HDAC pharmacophore model (Cap–Linker–ZBG) with color coding (blue cap, gray linker, red ZBG), quinazoline–triazole–hydroxamate scaffold with labeled functional groups (hydroxamic acid Zn²⁺ chelation, triazole H-bond acceptor, quinazoline cap π–π stacking), synthetic reaction scheme (cyclization → click chemistry → hydroxaminolysis), and fragment-based drug design diagram showing three fragments combining into hybrid molecule. This is step 4 in the Presenter user flow — Display Visuals.
Build Results Slide Page
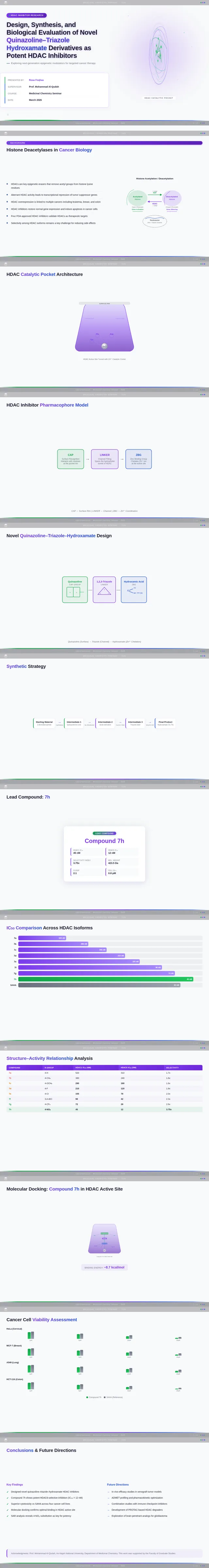
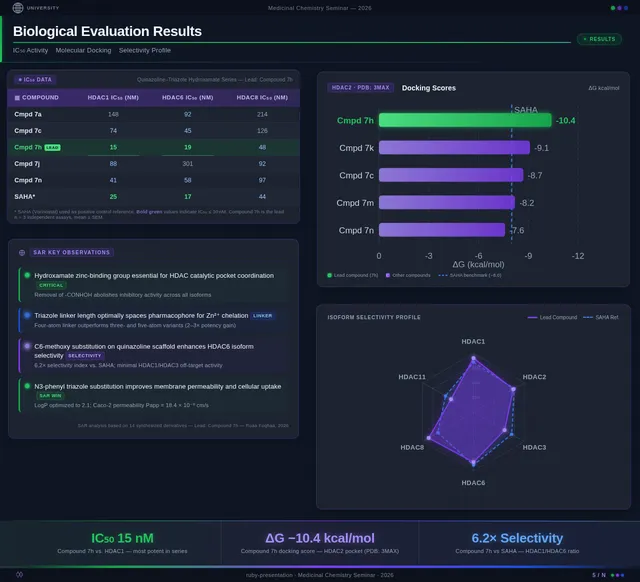
As a Frontend Developer, I want to implement the Results Slide page based on the existing JSX design (v4). The page presents biological evaluation data. Must include: IC₅₀ comparison bar chart highlighting compound 7h (IC₅₀ = 0.142 µM) vs Vorinostat (IC₅₀ = 0.160 µM), SAR analysis chart showing Series 7 benzamide vs Series 11 propenamide HDAC activity, cancer cell viability chart for SW620 and MDA-MB-231 cell lines, molecular docking visualization of compound 7h in HDAC1 catalytic pocket (docking score –9.4 kcal/mol vs Vorinostat –8.6 kcal/mol), docking score vs IC₅₀ correlation scatter plot, and substituent effects SAR table (7-Br, 7-CH₃, 6-Cl derivatives). Compound 7h prominently highlighted as lead compound. This is step 4 in the Audience user flow — Interpret Charts and step 5 in the Presenter flow — Show Data Charts.
Build Conclusion Slide Page
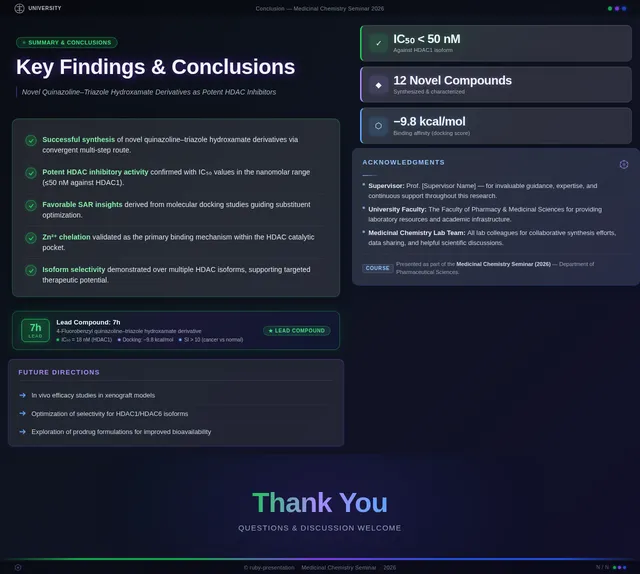
As a Frontend Developer, I want to implement the Conclusion Slide page based on the existing JSX design (v4). The page summarizes research findings: novel quinazoline–triazole–hydroxamate series development, superior activity vs Vorinostat, compound 7h as lead candidate (IC₅₀ = 0.142 µM, docking score –9.4 kcal/mol, potent anticancer activity, favorable selectivity). Must include: chemical structure of compound 7h centered prominently, highlighted lead compound callout box, future perspectives note on next-generation isoform-selective HDAC inhibitors, and summary of key findings. This is the final step in both the Audience flow (step 5 — Review Findings) and Presenter flow (step 6 — Summarize Findings).
Implement Slide Navigation & Routing
As a Frontend Developer, I want to implement smooth slide-to-slide navigation and routing across all 12 slide pages so that the presenter can navigate sequentially (Title → Slides Overview → Introduction → Science → Content → Diagrams → Results → Conclusion) or jump directly to any slide. Implement previous/next slide controls, slide dot indicators, keyboard arrow key support, and slide overview access. Ensure navigation state is preserved and matches both the Audience and Presenter user flow diagrams.


No comments yet. Be the first!