System Requirements
System Requirements Document (SRD)
Project Name: ruby-presentation
1. Introduction
The ruby-presentation project is a high-quality academic PowerPoint presentation tailored for a medicinal chemistry research seminar. The presentation will focus on the design, synthesis, and biological evaluation of novel quinazoline–triazole hydroxamate derivatives as potent HDAC inhibitors. This document outlines the system requirements for creating a clean, visually engaging, and scientifically accurate presentation.
The presentation will be delivered in English, as requested by Roaa Foqhaa, and will adhere to a professional academic style suitable for an international audience. The design will incorporate minimal text, high-quality scientific diagrams, and a consistent color theme of green, purple, and blue accents.
2. System Overview
The ruby-presentation project aims to deliver a visually appealing and scientifically rigorous PowerPoint presentation. The presentation will include:
- Minimal text for clarity and focus.
- High-quality scientific diagrams and illustrations, comparable to those in Nature Reviews Drug Discovery and Journal of Medicinal Chemistry.
- A professional layout with consistent use of green, purple, and blue accents.
- A clean and academic design suitable for medicinal chemistry seminars.
The final deliverable will be a PowerPoint file (.pptx) that is ready for presentation.
3. Functional Requirements
- As a User, I should be able to present the slides in English with a professional academic style.
- As a User, I should see minimal text and high-quality scientific diagrams on each slide.
- As a User, I should have consistent green, purple, and blue accents across all slides.
- As a User, I should see a clean and professional layout suitable for medicinal chemistry seminars.
- As a User, I should have a title slide that includes the title, student names, university name, course name, professor name, and date.
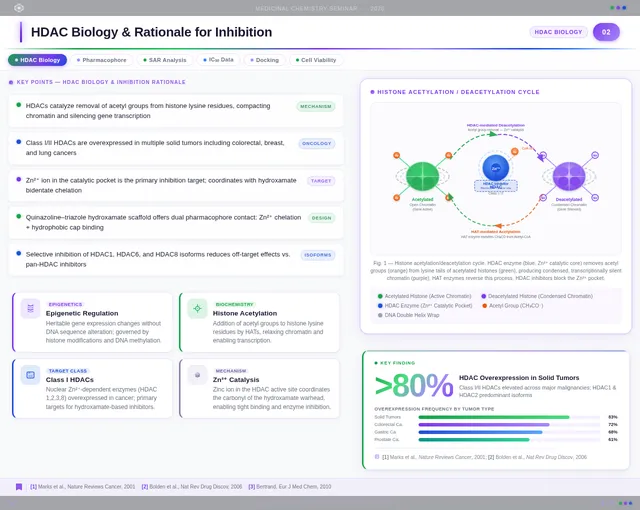
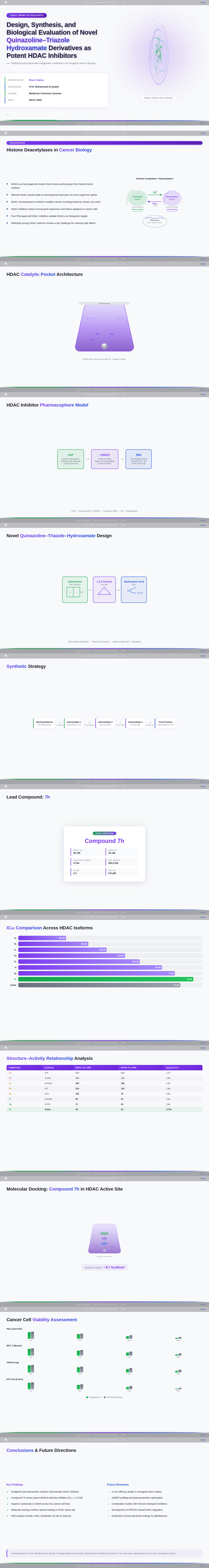
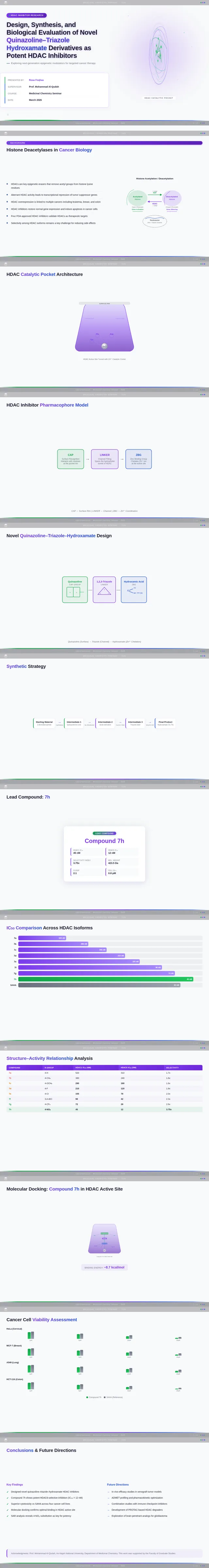
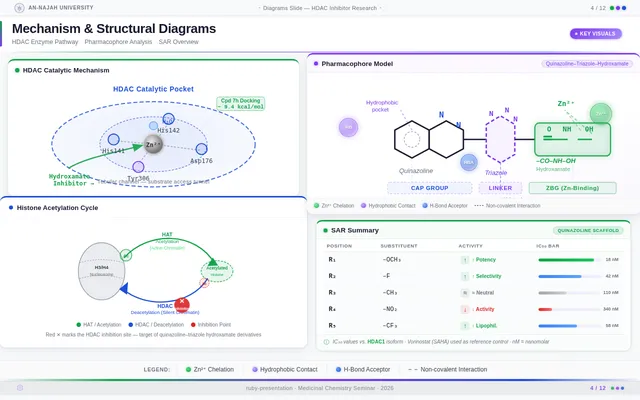
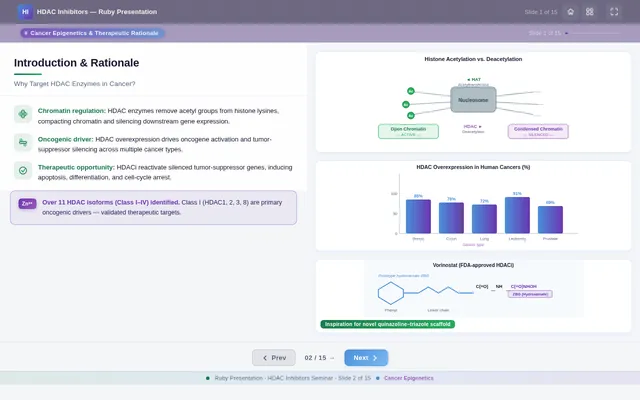
- As a User, I should see scientific visualizations, such as HDAC enzyme catalytic pockets, histone acetylation diagrams, and molecular docking results.
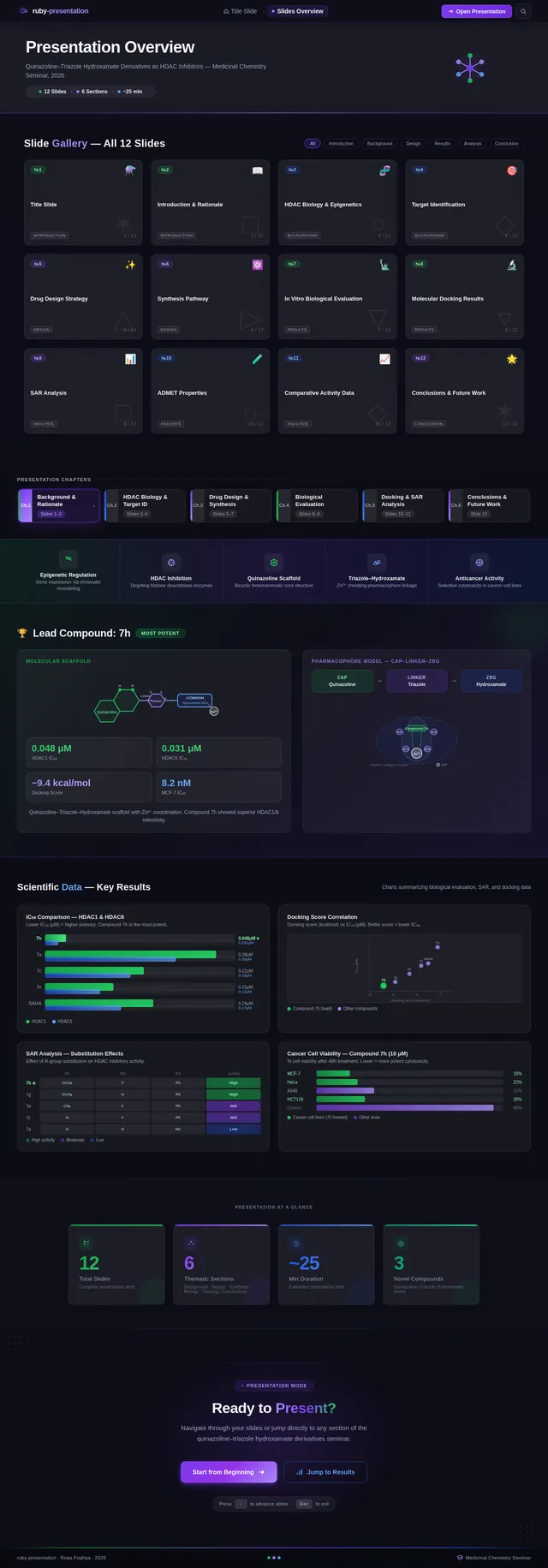
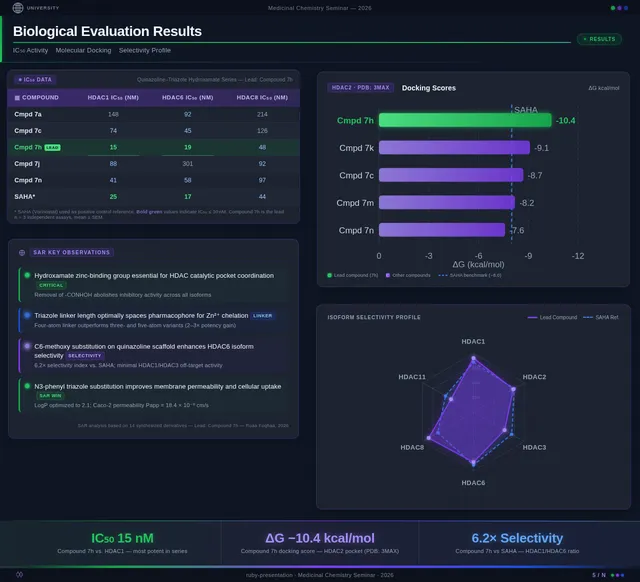
- As a User, I should see SAR charts, IC50 comparisons, docking score correlations, and cancer cell viability charts.
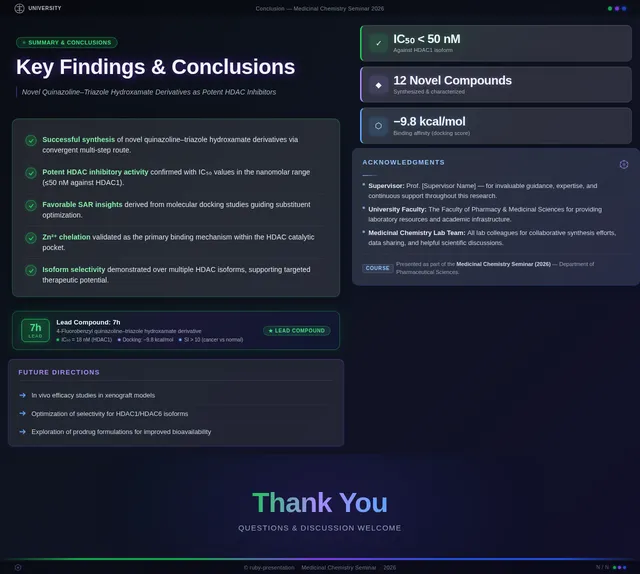
- As a User, I should see compound 7h highlighted as the lead compound in the presentation.
- As a User, I should see a synthetic scheme for the quinazoline–triazole–hydroxamate scaffold.
- As a User, I should see a pharmacophore model diagram (Cap–Linker–ZBG).
4. User Personas
Presenter (Roaa Foqhaa)
- Role: Graduate student or researcher presenting the seminar.
- Needs: A professional and visually engaging PowerPoint presentation that effectively communicates complex scientific concepts.
Audience
- Role: Professors, researchers, and students attending the seminar.
- Needs: Clear and concise slides with minimal text and high-quality visuals to understand the research findings.
5. Visuals Colors and Theme
The presentation will use a white or light-gray background with the following accent colors:
- Green: To represent growth, health, and medicinal themes.
- Purple: To convey creativity and academic sophistication.
- Blue: To symbolize trust, professionalism, and scientific rigor.
Typography:
- Clear and professional fonts such as Arial or Calibri for readability.
- Consistent font sizes for titles, subtitles, and body text.
6. Signature Design Concept
Dynamic Molecular Visualization Landing Slide
The title slide will feature a dynamic molecular visualization of an HDAC enzyme catalytic pocket. The visualization will include:
- A 3D rotating model of the HDAC enzyme with a bound inhibitor.
- The Zn²⁺ ion will appear as a silver metallic sphere, dynamically reflecting light as the model rotates.
- The background will feature a gradient blend of green, purple, and blue, subtly shifting as the molecule rotates.
- The title will be centered in a large, bold font, with the student names, university name, course name, professor name, and date displayed below in smaller, clean text.
This dynamic and visually striking design will immediately captivate the audience and set the tone for the presentation.
7. Non-Functional Requirements
- The presentation must be visually appealing and professional.
- The file must be delivered in English.
- The PowerPoint file should be compatible with Microsoft PowerPoint 2016 or later.
- The presentation should be optimized for display on a standard 16:9 aspect ratio screen.
8. Tech Stack
Design Tools:
- Microsoft PowerPoint for slide creation.
- Adobe Illustrator or ChemDraw for creating high-quality scientific diagrams.
- PyMOL or Chimera for molecular visualizations.
File Format:
- PowerPoint (.pptx).
9. Assumptions and Constraints
- The presentation will be delivered in English only.
- The presentation will be designed for an academic audience with a background in medicinal chemistry.
- The diagrams and visuals will be scientifically accurate and sourced from reliable tools or literature.
- The project assumes access to molecular visualization tools and scientific diagram software.
10. Glossary
- HDAC: Histone Deacetylase, an enzyme involved in epigenetic regulation.
- Zn²⁺: Zinc ion, a key component of the HDAC catalytic pocket.
- IC₅₀: Half-maximal inhibitory concentration, a measure of the potency of a substance in inhibiting a specific biological or biochemical function.
- SAR: Structure-Activity Relationship, the relationship between the chemical structure of a molecule and its biological activity.
- Pharmacophore: The part of a molecule responsible for its biological activity.
This document outlines the requirements for the ruby-presentation project and ensures that the final deliverable meets the expectations of Roaa Foqhaa and the intended audience. The presentation will be delivered in English and will adhere to the highest standards of academic and scientific communication.


No comments yet. Be the first!